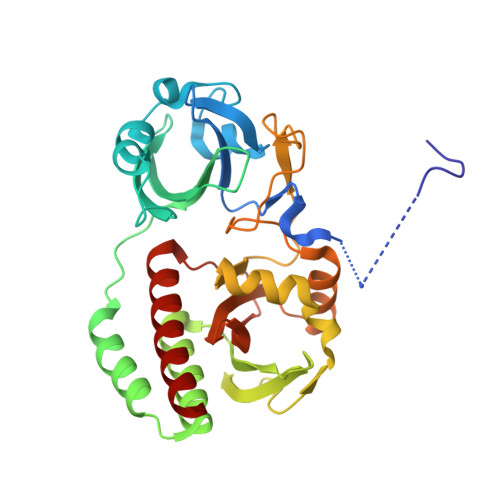
Designing brighter near-infrared fluorescent proteins: insights from structural and biochemical studies.
Baloban, M., Shcherbakova, D.M., Pletnev, S., Pletnev, V.Z., Lagarias, J.C., Verkhusha, V.V.(2017) Chem Sci 8: 4546-4557
- PubMed: 28936332
- DOI: https://doi.org/10.1039/c7sc00855d
- Primary Citation of Related Structures:
5VIK, 5VIQ, 5VIV - PubMed Abstract:
Brighter near-infrared (NIR) fluorescent proteins (FPs) are required for multicolor microscopy and deep-tissue imaging. Here, we present structural and biochemical analyses of three monomeric, spectrally distinct phytochrome-based NIR FPs, termed miRFPs. The miRFPs are closely related and differ by only a few amino acids, which define their molecular brightness, brightness in mammalian cells, and spectral properties. We have identified the residues responsible for the spectral red-shift, revealed a new chromophore bound simultaneously to two cysteine residues in the PAS and GAF domains in blue-shifted NIR FPs, and uncovered the importance of amino acid residues in the N-terminus of NIR FPs for their molecular and cellular brightness. The novel chromophore covalently links the N-terminus of NIR FPs with their C-terminal GAF domain, forming a topologically closed knot in the structure, and also contributes to the increased brightness. Based on our studies, we suggest a strategy to develop spectrally distinct NIR FPs with enhanced brightness.
Organizational Affiliation:
Department of Anatomy and Structural Biology and Gruss-Lipper Biophotonics Center , Albert Einstein College of Medicine , Bronx , New York 10461 , USA . Email: [email protected].