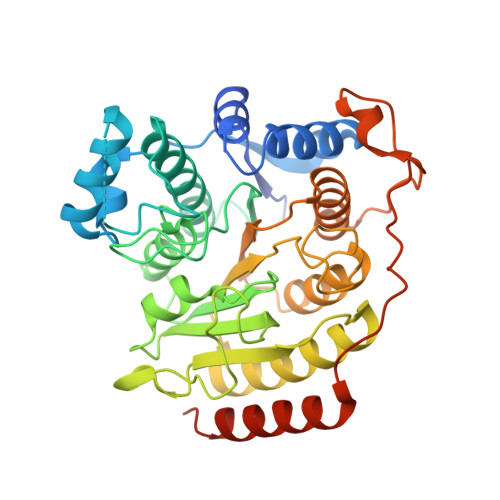
Biochemical and Structural Characterization of HDAC8 Mutants Associated with Cornelia de Lange Syndrome Spectrum Disorders.
Decroos, C., Christianson, N.H., Gullett, L.E., Bowman, C.M., Christianson, K.E., Deardorff, M.A., Christianson, D.W.(2015) Biochemistry 54: 6501-6513
- PubMed: 26463496
- DOI: https://doi.org/10.1021/acs.biochem.5b00881
- Primary Citation of Related Structures:
5D1B, 5D1C, 5D1D - PubMed Abstract:
Cornelia de Lange Syndrome (CdLS) spectrum disorders are characterized by multiple organ system congenital anomalies that result from mutations in genes encoding core cohesin proteins SMC1A, SMC3, and RAD21, or proteins that regulate cohesin function such as NIPBL and HDAC8. HDAC8 is the Zn(2+)-dependent SMC3 deacetylase required for cohesin recycling during the cell cycle, and 17 different HDAC8 mutants have been identified to date in children diagnosed with CdLS. As part of our continuing studies focusing on aberrant HDAC8 function in CdLS, we now report the preparation and biophysical evaluation of five human HDAC8 mutants: P91L, G117E, H180R, D233G, and G304R. Additionally, the double mutants D233G-Y306F and P91L-Y306F were prepared to enable cocrystallization of intact enzyme-substrate complexes. X-ray crystal structures of G117E, P91L-Y306F, and D233G-Y306F HDAC8 mutants reveal that each CdLS mutation causes structural changes that compromise catalysis and/or thermostability. For example, the D233G mutation disrupts the D233-K202-S276 hydrogen bond network, which stabilizes key tertiary structure interactions, thereby significantly compromising thermostability. Molecular dynamics simulations of H180R and G304R HDAC8 mutants suggest that the bulky arginine side chain of each mutant protrudes into the substrate binding site and also causes active site residue Y306 to fluctuate away from the position required for substrate activation and catalysis. Significantly, the catalytic activities of most mutants can be partially or fully rescued by the activator N-(phenylcarbamothioyl)-benzamide, suggesting that HDAC8 activators may serve as possible leads in the therapeutic management of CdLS.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania , Philadelphia, Pennsylvania 19104-6323, United States.