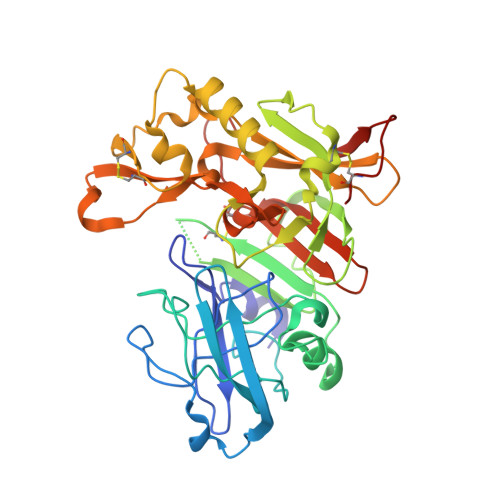
New aminoimidazoles as beta-secretase (BACE-1) inhibitors showing amyloid-beta (A beta ) lowering in brain.
Gravenfors, Y., Viklund, J., Blid, J., Ginman, T., Karlstrom, S., Kihlstrom, J., Kolmodin, K., Lindstrom, J., von Berg, S., von Kieseritzky, F., Bogar, K., Slivo, C., Swahn, B.M., Olsson, L.L., Johansson, P., Eketjall, S., Falting, J., Jeppsson, F., Stromberg, K., Janson, J., Rahm, F.(2012) J Med Chem 55: 9297-9311
- PubMed: 23017051
- DOI: https://doi.org/10.1021/jm300991n
- Primary Citation of Related Structures:
4B1C, 4B1D, 4B1E - PubMed Abstract:
Amino-2H-imidazoles are described as a new class of BACE-1 inhibitors for the treatment of Alzheimer's disease. Synthetic methods, crystal structures, and structure-activity relationships for target activity, permeability, and hERG activity are reported and discussed. Compound (S)-1m was one of the most promising compounds in this report, with high potency in the cellular assay and a good overall profile. When guinea pigs were treated with compound (S)-1m, a concentration and time dependent decrease in Aβ40 and Aβ42 levels in plasma, brain, and CSF was observed. The maximum reduction of brain Aβ was 40-50%, 1.5 h after oral dosing (100 μmol/kg). The results presented highlight the potential of this new class of BACE-1 inhibitors with good target potency and with low effect on hERG, in combination with a fair CNS exposure in vivo.
Organizational Affiliation:
Department of Medicinal Chemistry, AstraZeneca R&D Södertälje, SE-151 85 Södertälje, Sweden. [email protected]