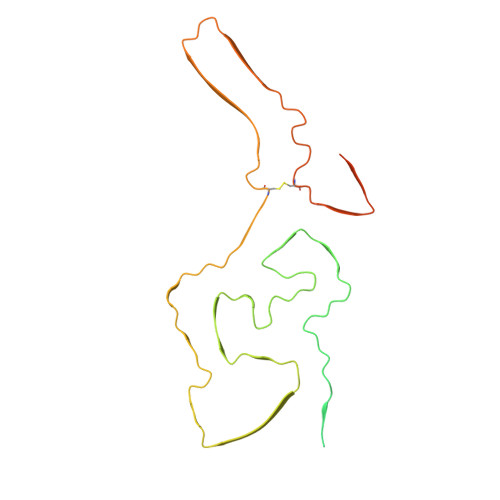
Cryo-EM structure of a natural prion: chronic wasting disease fibrils from deer.
Alam, P., Hoyt, F., Artikis, E., Soukup, J., Hughson, A.G., Schwartz, C.L., Barbian, K., Miller, M.W., Race, B., Caughey, B.(2024) Acta Neuropathol 148: 56-56
- PubMed: 39448454
- DOI: https://doi.org/10.1007/s00401-024-02813-y
- Primary Citation of Related Structures:
9DMY, 9DMZ - PubMed Abstract:
Chronic wasting disease (CWD) is a widely distributed prion disease of cervids with implications for wildlife conservation and also for human and livestock health. The structures of infectious prions that cause CWD and other natural prion diseases of mammalian hosts have been poorly understood. Here we report a 2.8 Å resolution cryogenic electron microscopy-based structure of CWD prion fibrils from the brain of a naturally infected white-tailed deer expressing the most common wild-type PrP sequence. Like recently solved rodent-adapted scrapie prion fibrils, our atomic model of CWD fibrils contains single stacks of PrP molecules forming parallel in-register intermolecular β-sheets and intervening loops comprising major N- and C-terminal lobes within the fibril cross-section. However, CWD fibrils from a natural cervid host differ markedly from the rodent structures in many other features, including a ~ 180° twist in the relative orientation of the lobes. This CWD structure suggests mechanisms underlying the apparent CWD transmission barrier to humans and should facilitate more rational approaches to the development of CWD vaccines and therapeutics.
Organizational Affiliation:
Laboratory of Neurological Infections and Immunity, Rocky Mountain Laboratories, Division of Intramural Research, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Hamilton, MT, 59840, USA.