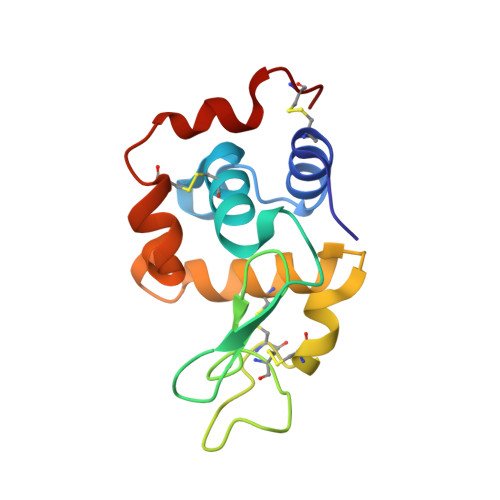
Structural studies of the effect that dimethyl sulfoxide (DMSO) has on cisplatin and carboplatin binding to histidine in a protein.
Tanley, S.W., Schreurs, A.M., Kroon-Batenburg, L.M., Meredith, J., Prendergast, R., Walsh, D., Bryant, P., Levy, C., Helliwell, J.R.(2012) Acta Crystallogr D Biol Crystallogr 68: 601-612
- PubMed: 22525758
- DOI: https://doi.org/10.1107/S0907444912006907
- Primary Citation of Related Structures:
4DD0, 4DD1, 4DD2, 4DD3, 4DD7, 4DD9, 4DDA, 4DDB, 4DDC - PubMed Abstract:
The anticancer complexes cisplatin and carboplatin target the DNA major groove, forming intrastrand and interstrand cross-links between guanine bases through their N7 atoms, causing distortion of the DNA helix and apoptotic cell death. A major side effect of these drugs is toxicity, which is caused via binding to many proteins in the body. A range of crystallographic studies have been carried out involving the cocrystallization of hen egg-white lysozyme (HEWL) as a test protein with cisplatin and carboplatin in aqueous and dimethyl sulfoxide (DMSO) conditions. Different cryoprotectants, glycerol and Paratone, were used for each of the cisplatin and carboplatin cocrystallization cases, while silicone oil was used for studies involving N-acetylglucosamine (NAG). Both cisplatin and carboplatin do not bind to HEWL in aqueous media on the timescales of the conditions used here, but upon addition of DMSO two molecules of cisplatin or carboplatin bind either side of His15, which is the only His residue in lysozyme and is assumed to be an imidazolyl anion or a chemical resonance moiety, i.e. both imidazole N atoms are chemically reactive. To identify the platinum-peak positions in the 'with DMSO conditions', anomalous scattering maps were calculated as a cross-check with the F(o) - F(c) OMIT maps. Platinum-occupancy σ values were established using three different software programs in each case. The use of EVAL15 to process all of the diffraction data sets provided a consistent platform for a large ensemble of data sets for the various protein and platinum-compound model refinements with REFMAC5 and then SHELXTL. Overall, this extensive set of crystallization and cryoprotectant conditions allowed a systematic evaluation of cisplatin and carboplatin binding to lysozyme as a test protein via detailed X-ray crystal structure characterizations. DMSO is used as a super-solvent for drug delivery as it is deemed to cause no effect upon drug binding. However, these results show that addition of DMSO causes the platinum anticancer drugs to bind to HEWL. This effect should be considered in toxicity assessments of these drugs and perhaps more widely.
Organizational Affiliation:
School of Chemistry, Faculty of Engineering and Physical Sciences, University of Manchester, Brunswick Street, Manchester M13 9PL, England.